Key Takeaways
- Genotyping vs. Sequencing: Genotyping looks at specific, pre-selected DNA positions, while sequencing reads entire sections of DNA for more comprehensive data.
- Targeted Gene Panels: These are used to screen for specific known disease-causing variants, such as those linked to breast cancer or epilepsy.
- Exome Sequencing: Focuses on the 2% of the genome that codes for proteins, offering a balance between cost and diagnostic power.
- Whole Genome Sequencing (WGS): The most thorough method, reading nearly all 6.4 billion bases to provide a complete genetic picture.
- Emerging Tech: Long-read sequencing is improving the detection of complex DNA deletions and duplications that short-read methods might miss.
What are the main types of DNA testing?
There are two main types of DNA testing that are used in research and in direct-to-consumer tests: genotyping and sequencing.

- Genotyping: Tests 500,000–1,000,000 specific positions known to vary between people. It is cost-effective but limited to pre-selected data points.
- Sequencing: Reads full stretches of DNA without needing to pre-select positions. It provides significantly more data but at a higher cost.
Within these two broad categories, there are many distinct variants of test, each designed for different purposes and coverage profiles. For example, there are hundreds of different genotype arrays – each array covers different positions in the genome and may be used for slightly different purposes. An example of this is the Illumina Global Screening array, which is very popular with direct-to-consumer genetic testing companies; this genotype array covers around 640,000 positions. Another example is the UKBiobank Array, which was used to test 500,000 participants in the UK biobank (as well as many other research projects) and covers 820,000 positions.
Types of DNA sequencing
Within the category of sequencing, there are three main tests that are used for research and diagnostic testing, as well as increasingly in direct-to-consumer testing.
Targeted gene panels
Targeted gene panels involve pre-selecting a set of sequences (usually protein-coding genes) to test in order to look for known disease-causing genetic variants. Targeted Gene Panels are a two step process where DNA is first filtered to capture only the pre-selected set of sequences or "targets." Then the captured DNA is sent off to be sequenced.
For example, a person with a family history of breast cancer may be tested for a panel of a few dozen genes that are known to cause breast cancer. As another example, a child or young adult who has unexplained epilepsy/seizures would be tested for a completely different set of genes known to cause epilepsy. There are hundreds of genes that are known to cause epilepsy when their sequence is changed, and an epilepsy gene panel would tests all of them at once.
For diagnosing rare and chronic disease, gene panels can often find the genetic variant causing the condition, but sometimes result in a "diagnostic odyssey" where nothing is found in the panel, and further testing needs to be done, sometimes taking years to find the cause of disease. For this reason, panel sequencing has started to be replaced by exome sequencing for many genetic disorders.
Exome sequencing

Exome sequencing involves a two-step process, similar to panel sequencing, where the DNA is filtered to capture only DNA that maps to one of the 20,000 protein-coding genes in the genome. The second step is to sequence the DNA that maps to a protein coding gene, which corresponds to ~2% of the total DNA.
Exome sequencing is more expensive than targeted panels, but offers a materially broader diagnostic yield—particularly for patients with complex or previously unresolved presentations. In precision medicine trials, exome sequencing can also support exploratory endpoints and cohort characterisation beyond the primary eligibility variant, providing additional data on variant burden and co-occurring mutations relevant to stratification or response analysis.
However, the main drawback of exome sequencing is that it misses 98% of the DNA. This can be the cause of missed diagnoses in rare or chronic disorders, when the mutation is not in the ~2% of the DNA that codes for proteins. Exome sequencing can also make it more challenging to detect deletions or duplications of DNA. For both of these reasons, many research projects and diagnostic sequencing efforts such as the 100,000 genomes project, are now turning to whole genome sequencing.
Whole genome sequencing
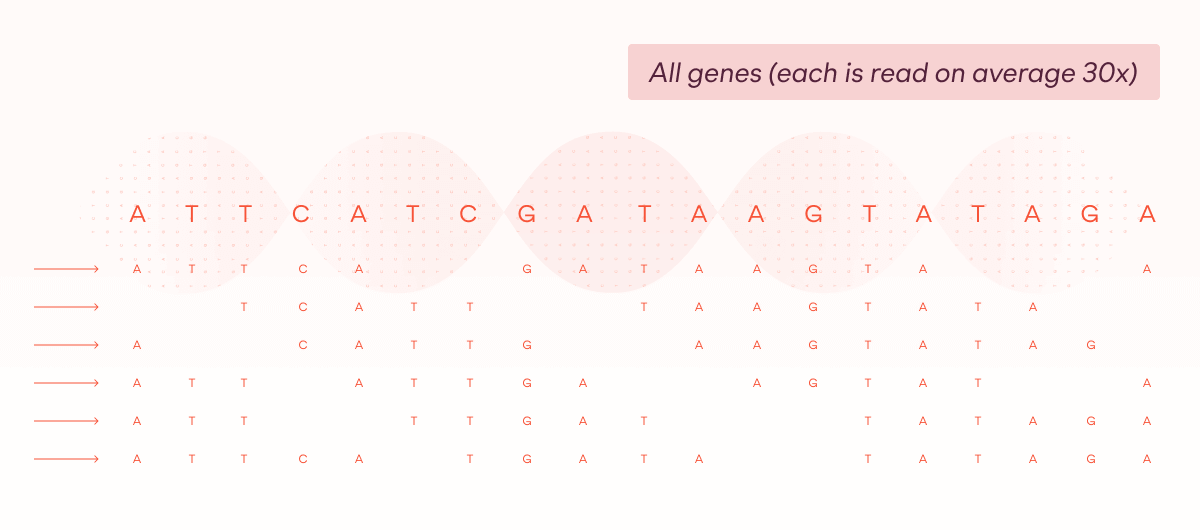
Whole genome sequencing does not require any filtering step – instead, the DNA is sent straight for sequencing, and there are on average approximately 30 reads of every position of the DNA. This produces a high-fidelity representation of the DNA sequence across nearly all 6.4 billion bases.
The majority of whole genome sequencing today uses "short read" technology – this means that about 100-150 bases of DNA are read at a time. As these reads overlap each other, they can be stitched together to form a nearly complete picture.
Other sequencing approaches
Besides the approaches covered above, there are others you may have heard of, specifically "low pass" whole genome sequencing and long read sequencing:
Low pass sequencing is a form of whole genome sequencing, but on average every base is covered by less than one sequencing read. In theory, statistical tools can be applied to piece together the missing sequence, making the data comparable to genotyping arrays, but with no need to pre-select positions to test. However, it is in no way a replacement for whole genome sequencing or exome sequencing.
Long read sequencing represents a technically significant advance in whole genome sequencing that reads very long stretches of DNA – sometimes tens of thousands of bases at a time, in contrast with the more commonly used short read sequencing. One big advantage of long read sequencing is that it can help to better detect deletions and duplications in DNA, which can be the cause of severe genetic conditions, as well as more subtle variations in traits or disease risk.
Another advantage of long read sequencing is that some technologies get information about methylation (the modifications to the DNA bases, sometimes referred to as ‘epigenetics’) at the same time as the DNA. Long read whole genome sequencing remains cost-prohibitive for most clinical programs at present, but as costs decline, its ability to resolve structural variants and capture epigenetic data simultaneously makes it increasingly relevant to studies where standard short-read methods have demonstrated eligibility confirmation gaps.


